This page provides resources for:
- TUTORIALS: video tutorials that demonstrate various aspects of using the code base.
- HOW-TO-GUIDES: step-by-step user guidelines for using DeepLabCut on your own datasets (or on demo data)
- EXPLANATIONS: resources on understanding how DeepLabCut works (WIP)
- REFERENCES: read the science behind DeepLabCut
-
a set of videos that span the types of behaviors you want to track. Having 10 videos that include different backgrounds, different individuals, and different postures is MUCH better than 1 or 2 videos of 1 or 2 different indivduals (i.e. 10-20 frames from each of 10 videos is much better than 50-100 frames from 2 videos).
-
a computer. If you want to use DeepLabCut on your own computer for many experiments, then you should get an NVIDIA GPU. See technical specs here. In the short term, or for testing out our software you can run short demos on COLAB (see how), or on your CPU.
-
no specific cameras are required; color, monochrome, etc is all fine. If you can see what you want to measure, then this will work for you (given enough labeled-data).
-
no specific computer is required (but see recommendations above), our software works on Linux, Windows, and MacOS, although we recommend Ubuntu.
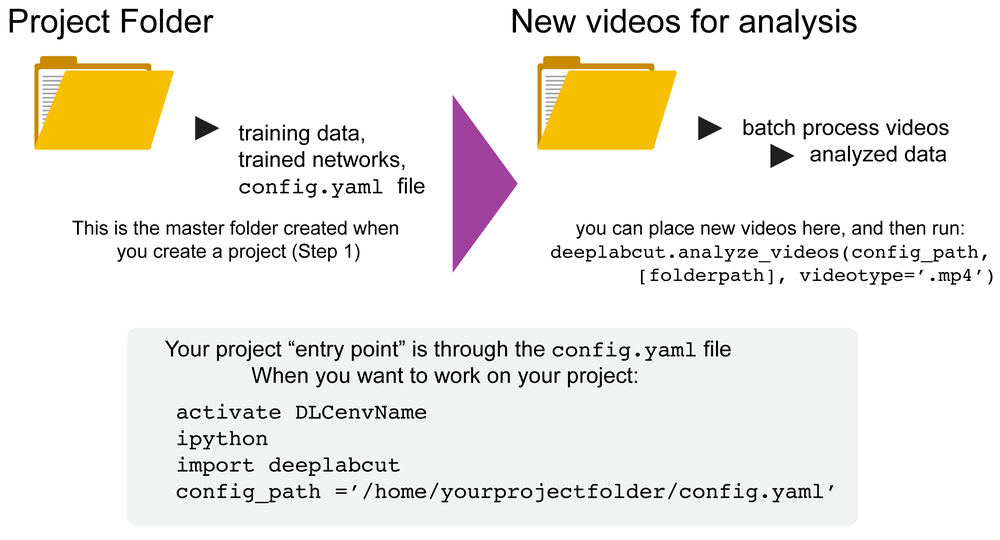
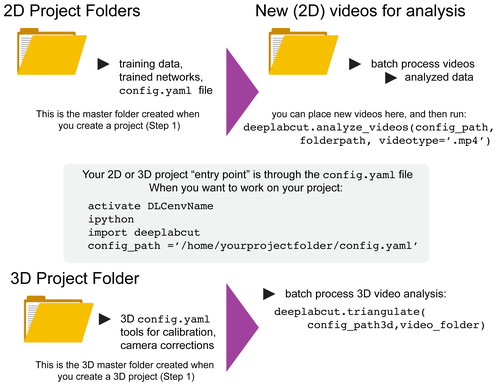
Overview: DeepLabCut is a software package for markerless pose estimation of animals performing various tasks. The software can manage multiple projects for various tasks. Each project is identified by the name of the project (e.g. TheBehavior), name of the experimenter (e.g. YourName), as well as the date at creation. This project folder holds a config.yaml (a text document) file containing various (project) parameters as well as links the data of the project.
You can have as many projects on your computer as you wish. You can have DeepLabCut installed in an environment and always exit and return to this environment to run the code. You just need to point to the correct config.yaml file to jump back in! The documentation below will take you through the individual steps.
Coming soon: as of 2.2 we support multi-animal projects. The workflow is highly similar, but with some key steps that differ. Please carefully review the functions below for more details. You can search/look for maDeepLabCut for specific steps that are changed. But until the package (2.2) is released, these steps are not functional.
As of 2.1+ we support a full Project Manager GUI to run the software. As of 2.0.7+ we support 3D analysis directly inside our package. Please see the 3D details here.
This page contains a list of the essential functions of DeepLabCut as well as demos. There are many optional parameters with each described function, which you can find here. For additional assistance, you can use the help function to better understand what each function does.
Great - now that you get the overall workflow, let's jump in. Here, you have several options. Option 1 is good for a quick demo on our data. Option 2 is the perfect place for beginners who want to start using DeepLabCut on your own data. Option 3 is best for more advanced users, as with the terminal interface you get the most versatility and options.
We provide Jupyter and COLAB notebooks for using DeepLabCut on both a pre-labeled dataset, and on the end user’s own dataset. See all the demo's here! Please note that GUIs are not easily supported in Jupyter in MacOS, as you need a framework build of python. While it's possible to launch them with a few tweaks, we recommend using the Project Manager GUI or terminal, so please follow the instructions below.
Start iPython, or if you are using MacOS, you must use pythonw vs. typing ipython or python, but otherwise it's the same.
If you are using DeepLabCut on the cloud, you cannot use the GUIs and you need to first set DLClight=True. Please read more here, and in our Protocol paper here.
Open an ipython session, import the package, and launch by typing in the terminal:
ipython
import deeplabcut
deeplabcut.launch_dlc()Or, simply open the terminal and type:
python -m deeplabcutThat's it! Follow the GUI for details
*please note, we provide a quick-guide of the commands at the bottom of this page.
Also, if you are using MacOS, you must use pythonw vs. typing ipython, but otherwise it's the same.
If you are using DeepLabCut on the cloud, you cannot use the GUIs and you need to first set DLClight=True. Please read more here, and in our Protocol paper here.
Open an ipython session and import the package by typing in the terminal:
ipython
import deeplabcutTIP: for every function there is a associated help document that can be viewed by adding a ? after the function name; i.e. deeplabcut.create_new_project?. To exit this help screen, type :q.
deeplabcut.create_new_project('ProjectName','YourName', ['/usr/FullPath/OfVideo1.avi','/usr/FullPath/OfVideo2.avi','/usr/FullPath/OfVideo1.avi'], copy_videos = True/False)Tip: if you want to place the project folder somewhere please pass : working_directory = 'FullPathOftheworkingDirectory'
maDeepLabCut: As of 2.2 when you create a project also pass: multianimal=True
- Note, if you are a Ubuntu user the path should look like:
['/home/username/yourFolder/video1.mp4']; if you are a Windows user, it should look like:[r'C:\username\yourFolder\video1.mp4'] - Note, you can also put
config_path =in front of the above line to create the path to the config.yaml that is used in the next step, i.e.config_path=deeplabcut.create_project(...))- If you do not, we recommend setting a variable so this can be easily used! Once you run this step, the conig_path is printed for you once you run this line, so set a variable for ease of use, i.e. something like:
config_path = '/thefulloutputpath/config.yaml'- just be mindful of the formatting for Windows vs. Linux, see below.
(more details here)
- open the config.yaml file (in a text editor (like atom, gedit, vim etc.)), which can be found in the subfolder created when you set your project name, to change parameters and identify label names! This is a crucial step.
maDeepLabCut: As of 2.2 you also need to modify a few more parameters in the config.yaml file, see here.
(PLEASE see more details here)
- set the config_path (LINUX):
config_path = '/home/computername/DeepLabCut/yourprojectname/config.yaml'- set the config_path (WINDOWS):
config_path = r'C:\home\computername\DeepLabCut\yourprojectname\config.yaml'- or you already set it as a variable in step "create_new_project" directly above.
mini-demo: create project and edit the yaml file
deeplabcut.extract_frames(config_path, mode='automatic', algo='kmeans', crop = True/False)(more details here) *update: as of 2.0.5 (spring 2019) checkcropping=True is dropped; you now just the option to directly draw a rectangle over the image to crop before extraction (i.e. there no need to manually change in config.yaml then check).
deeplabcut.label_frames(config_path)maDeepLabCut: As of 2.2 there is a new multi-animal labeling GUI:
deeplabcut.label_frames(config_path, multianimal=True)(more details here)
mini-demo: using the GUI to label
deeplabcut.check_labels(config_path)(more details here)
deeplabcut.create_training_dataset(config_path)or to compare different neural networks use:
deeplabcut.create_training_model_comparision(config_path, num_shuffles=1, net_types=['resnet_50'], augmenter_types=['default', 'imgaug'] )(more details here)
deeplabcut.train_network(config_path)(more details here)
deeplabcut.evaluate_network(config_path, plotting = True)(more details here)
- Please note that novel videos DO NOT need to be added to the config.yaml file. You can simply have a folder elsewhere on your computer and pass the video folder (then it will analyze all videos of the specified type (i.e.
videotype='.mp4'), or pass the path to the folder or exact video(s) you wish to analyze:
deeplabcut.analyze_videos(config_path,[`/fullpath/project/videos/'], videotype='.mp4', save_as_csv = True)Here are some tips for scaling up your analysis: https://github.com/AlexEMG/DeepLabCut/wiki/Batch-Processing-your-Analysis
You can also filter the predicted bodyparts by:
deeplabcut.filterpredictions(config_path,['/fullpath/project/videos/reachingvideo1.avi'])Note, this creates a file with the ending filtered.h5 that you can use for further analysis. This filtering step has many parameters, so please see the full docstring by typing: deeplabcut.filterpredictions?
Create videos:
deeplabcut.create_labeled_video(config_path, [`/analysis/project/videos/reachingvideo1.avi','/fullpath/project/videos/reachingvideo2.avi'],filtered = True)Plot the outputs:
deeplabcut.plot_trajectories(config_path,['/fullpath/project/videos/reachingvideo1.avi'],filtered = True)(more details here)
deeplabcut.extract_outlier_frames(config_path,['full/videofile_path'])(more details here)
(refinement and augmentation of the training dataset)
deeplabcut.refine_labels(config_path)mini-demo: using the refinement GUI, a user can load the file then zoom, pan, and edit and/or remove points:
When done editing the labels, merge: PRO TIP: if you addded new data, even without refining, i.e. you added and labeled frames from new videos, also use merge before creating a new training data set!
deeplabcut.merge_datasets(config_path)Now, create a new training set and re-train (same steps as above)! i.e.
deeplabcut.create_training_dataset(config_path)
deeplabcut.train_network(config_path)(more details here)
In ipython/Jupyter notebook:
deeplabcut.nameofthefunction?
In python or pythonw:
help(deeplabcut.nameofthefunction)
You can always exit an conda environment and easily jump back into a project by simply:
Linux/MacOS formatting example:
source activate yourdeeplabcutEnvName
ipython or pythonw
import deeplabcut
config_path ='/home/yourprojectfolder/config.yaml'
Windows formatting example:
activate yourdeeplabcutEnvName
ipython
import deeplabcut
config_path = r'C:\home\yourprojectfolder\config.yaml'
Now, you can run any of the functions desribed in this documentation.
There is also helper code to further analyze your data or scale up analsis here: https://github.com/AlexEMG/DLCutils/
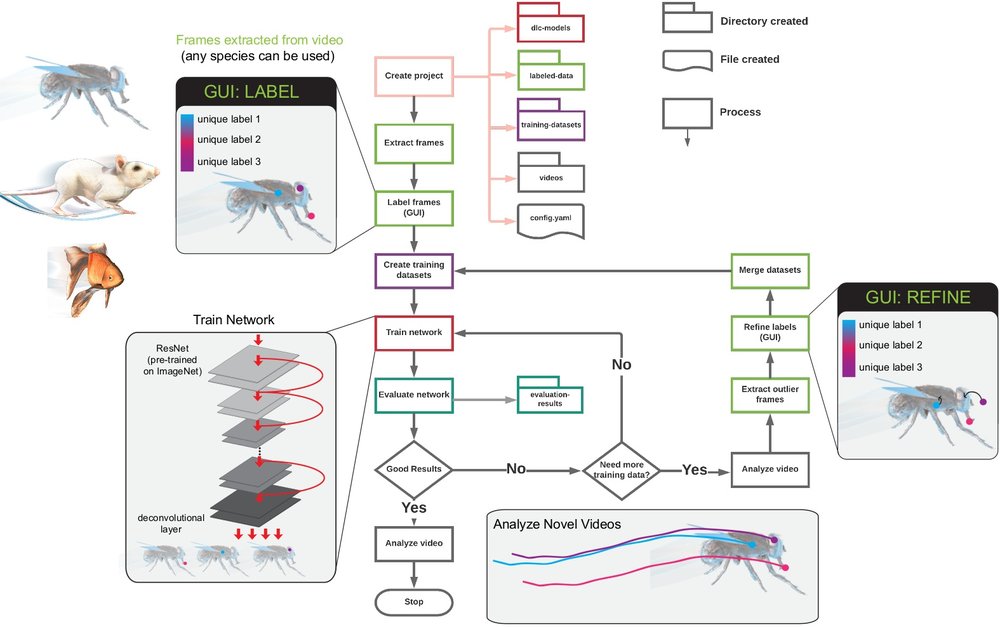
The 12 main steps to take you from project creation to analyzed videos:
Open ipython in the terminal:
import deeplabcut
Create a new project:
deeplabcut.create_new_project('project_name','experimenter’,['path of video 1','path of video2',..])
Set a config_path variable for ease of use:
config_path = 'yourdirectory/project_name/config.yaml'
Extract frames:
deeplabcut.extract_frames(config_path)
Label frames:
deeplabcut.label_frames(config_path)
Check labels [OPTIONAL]:
deeplabcut.check_labels(config_path)
Create training dataset:
deeplabcut.create_training_dataset(config_path)
Train the network:
deeplabcut.train_network(config_path)
Evaluate the trained network:
deeplabcut.evaluate_network(config_path)
Video analysis:
deeplabcut.analyze_videos(config_path, ['path of folder with videos'])
Plot results (trajectories):
deeplabcut.plot_trajectories(config_path, ['path of folder with videos'])
Create a video:
deeplabcut.create_labeled_video(config_path, ['path of folder with videos'])
Return to readme.