-
Notifications
You must be signed in to change notification settings - Fork 23
/
00-install_name_plot.Rmd
228 lines (146 loc) · 9.24 KB
/
00-install_name_plot.Rmd
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
148
149
150
151
152
153
154
155
156
157
158
159
160
161
162
163
164
165
166
167
168
169
170
171
172
173
174
175
176
177
178
179
180
181
182
183
184
185
186
187
188
189
190
191
192
193
194
195
196
197
198
199
200
201
202
203
204
205
206
207
208
209
210
211
212
213
214
215
216
217
218
219
220
221
222
223
224
225
226
227
---
title: 'Introduce Yourself'
author: "Alison Hill"
output:
html_document:
theme: flatly
toc: TRUE
toc_float: TRUE
toc_depth: 2
number_sections: TRUE
---
```{r setup, include = FALSE, cache = FALSE}
knitr::opts_chunk$set(error = TRUE, comment = NA, warnings = FALSE, errors = FALSE, messages = FALSE, tidy = FALSE)
```
```{r load-packages, include = FALSE}
library(tidyverse)
library(babynames)
```
> Read all the way through step 6, and note that there is a file that needs to be turned in to Sakai before Wednesday at noon!
# Overview
This guide will lead you through the steps to install and use [R, a free and open-source software environment for statistical computing and graphics.](https://www.r-project.org)
What is R?
* **R** is the name of the programming language itself, based off of S from Bell Labs, which users access through a command-line interpreter (`>`)
What is RStudio?
* **RStudio** is a powerful and convenient user interface that allows you to access the R programming language along with a lot of other bells and whistles that enhance functionality (and sanity).
Our end goal is to get you looking at a screen like this:

# Install R
Install R from [CRAN, the Comprehensive R Archive Network](https://cran.rstudio.com). Please choose a **precompiled binary distribution** for your operating system.
* If you need more help, check out one of the following videos (courtesy of Roger Peng at Johns Hopkins Biostatistics):
- [Installing R on a mac](https://www.youtube.com/watch?v=Icawuhf0Yqo&feature=youtu.be)
- [Installing R on windows](https://www.youtube.com/watch?v=mfGFv-iB724&feature=youtu.be)
* If you need even more help, read this [step-by-step guide](https://beckmw.files.wordpress.com/2014/09/r_install_guide.pdf), including screenshots.
## Check in
Launch R. You should see one console with a command line interpreter (`>`). Close R.
# Install RStudio
Install the free, open-source edition of RStudio: http://www.rstudio.com/products/rstudio/download/
RStudio provides a powerful user interface for R, called an *integrated development environment*. RStudio includes:
* a console (the standard command line interface: `>`),
* a syntax-highlighting editor that supports direct code execution, and
* tools for plotting, history, debugging and workspace management.
## Check in
Launch RStudio. You should get a window similar to the screenshot you see [here](https://www.rstudio.com/wp-content/uploads/2014/04/rstudio-workbench.png), but yours will be empty. Look at the bottom left pane: this is the same console window you saw when you opened R in step 1.15.
* Place your cursor where you see `>` and type `x <- 2 + 2`, hit enter or return, then type `x`, and hit enter/return again.
* If `[1] 4` prints to the screen, you have successfully installed R and RStudio, and you can move onto installing packages.
# Install packages
The version of R that you just downloaded is considered base R, which provides you with good but basic statistical computing and graphics powers. For analytical and graphical super-powers, you'll need to install add-on packages, which are user-written, to extend/expand your R capabilities. Packages can live in one of two places:
* They may be carefully curated by CRAN (which involves a thorough submission and review process), and thus are easy install using `install.packages("name_of_package", dependencies = TRUE)`.
* Alternatively, they may be available via GitHub. To download these packages, you first need to install the `devtools` package.
```{r devtools, eval = FALSE}
install.packages("devtools")
library(devtools)
install_github("name_of_package")
```
Place your cursor in the console again (where you last typed `x` and `[4]` printed on the screen). You can use the first method to install the following packages directly from CRAN, all of which we will use:
* [`dplyr`,](http://dplyr.tidyverse.org)
* [`ggplot2`,](http://ggplot2.tidyverse.org)
* [`babynames`](https://github.com/hadley/babynames)
You can download all of these at once, too:
```{r c_install, eval = FALSE}
install.packages(c("dplyr", "ggplot2", "babynames"),
dependencies = TRUE)
```
<p class="text-info"> __Heads up:__ We should formally introduce the combine command, [`c()`](http://stat.ethz.ch/R-manual/R-patched/library/base/html/c.html), used above. You will use this often- any time you want to combine things into a vector.</p>
```{r c_demo}
c("hello", "my", "name", "is", "alison")
c(1:3, 20, 50)
```
Mind your use of quotes carefully with packages.
* To *install* a package, you put the name of the package in quotes as in `install.packages("name_of_package")`.
* To *use* an already installed package, you must load it first, as in `library(name_of_package)`, leaving the name of the package bare. You only need to do this once per RStudio session.
* If you want *help*, no quotes are needed: `help(name_of_package)` or `?name_of_package`.
* If you want the *citation* for a package (and you should give credit where credit is due), ask R as in `citation("name_of_package")`.
```{r show_dplyr, eval = F}
install.packages("dplyr", dependencies = TRUE)
library(dplyr)
help("dplyr")
```
```{r help_dplyr}
citation("ggplot2")
```
<p class="text-info"> __Heads up:__ R is **case-sensitive**, so `?dplyr` works but `?Dplyr` will not. Likewise, a variable called `A` is different from `a`.</p>
# Make a name plot
Open a new R script in RStudio by going to `File --> New File --> R Script`. For this first foray into R, I'll give you the code, so sit back and relax and feel free to copy and paste my code with some small tweaks.
First load the packages:
```{r show_name_packages, eval = F}
library(babynames) # contains the actual data
library(dplyr) # for manipulating data
library(ggplot2) # for plotting data
```
Next, we'll follow [best practices for inspecting a freshly read dataset](https://cran.r-project.org/doc/contrib/de_Jonge+van_der_Loo-Introduction_to_data_cleaning_with_R.pdf). Also, see ["What I do when I get a new data set as told through tweets"](http://simplystatistics.org/2014/06/13/what-i-do-when-i-get-a-new-data-set-as-told-through-tweets/) for more ideas about exploring a new dataset. Here are some critical commands to obtain a high-level overview (HLO) of your freshly read dataset in R. We'll call it saying hello to your dataset:
```{r hlo_dataset}
glimpse(babynames) # dplyr
head(babynames) # base R
tail(babynames) # same
names(babynames) # same
```
If you have done the above and produced sane-looking output, you are ready for the next bit. Use the code below to create a new data frame called `alison`.
```{r manipulate_name_data}
alison <- babynames %>%
filter(name == "Alison" | name == "Allison") %>%
filter(sex == "F")
```
* The first bit makes a new dataset called `alison` that is a copy of the `babynames` dataset- the `%>%` tells you we are doing some other stuff to it later.
* The second bit `filters` our `babynames` to only keep rows where the `name` is either Alison or Allison (read `|` as _"or"_.)
* The third bit applies another `filter` to keep only those where sex is female.
Let's check out the data.
```{r}
alison
glimpse(alison)
```
Again, if you have sane-looking output here, move along to plotting the data!
```{r plot_name_data}
plot <- ggplot(alison, aes(x = year,
y = prop,
group = name,
color = name)) +
geom_line()
```
Now if you did this right, you will not see your plot! Because we saved the `ggplot` with a name (`plot`), R just saved the object for you. But check out the top right pane in RStudio again: under `Values` you should see `plot`, so it is there, you just have to ask for it. Here's how:
```{r}
plot
```
# Make a new name plot
Edit my code above to create a new dataset. Pick 2 names to compare how popular they each are (these could be different spellings of your own name, like I did, but you can choose any 2 names that are present in the dataset). Make the new plot, changing the name of the first argument `alison` in `ggplot()` to the name of your new dataset.
## Save and share
Save your work so you can share your favorite plot with us. You will not like the looks of your plot if you mouse over to `Export` and save it. Instead, use `ggplot2`'s command for saving a plot with sensible defaults:
```{r eval = FALSE}
help(ggsave)
```
```{r eval = F}
ggsave("alison_hill.pdf", plot) # please make the filename unique!
```
Upload this exported plot to Sakai before Wednesday at noon.
# Other cool `babynames` projects
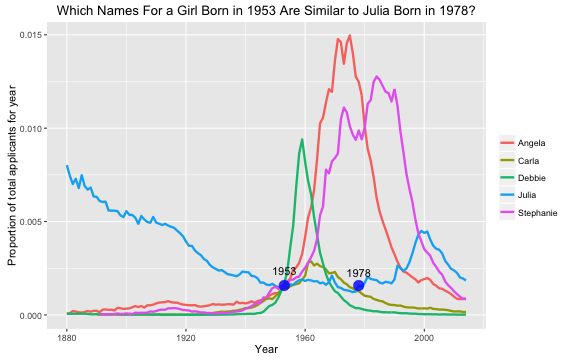
- Julia Silge *'My Baby Boomer Name Might Have Been "Debbie"'*: https://juliasilge.com/blog/my-baby-boomer-name/
- Use Julia's Shiny app: https://juliasilge.shinyapps.io/PredictNamesApp/
- Hilary Parker: Hilary: The Most Poisoned Baby Name in US History: https://hilaryparker.com/2013/01/30/hilary-the-most-poisoned-baby-name-in-us-history/

# Resources
- https://alison.rbind.io/html/jamboree_heart_ggplot.html
- http://moderndive.com/2-getting-started.html
- http://r4ds.had.co.nz
- https://www.rstudio.com/resources/cheatsheets/
- https://rweekly.org