-
Notifications
You must be signed in to change notification settings - Fork 44
Home
Molecular heterogeneities bring great challenges for cancer diagnosis and treatment. Recent advances in single cell RNA-sequencing (scRNA-seq) technology make it possible to study cancer transcriptomic heterogeneities at single cell level.
Here, we develop an R package named scCancer which focuses on processing and
analyzing scRNA-seq data for cancer research. Except basic data processing steps,
this package takes several special considerations for cancer-specific features.
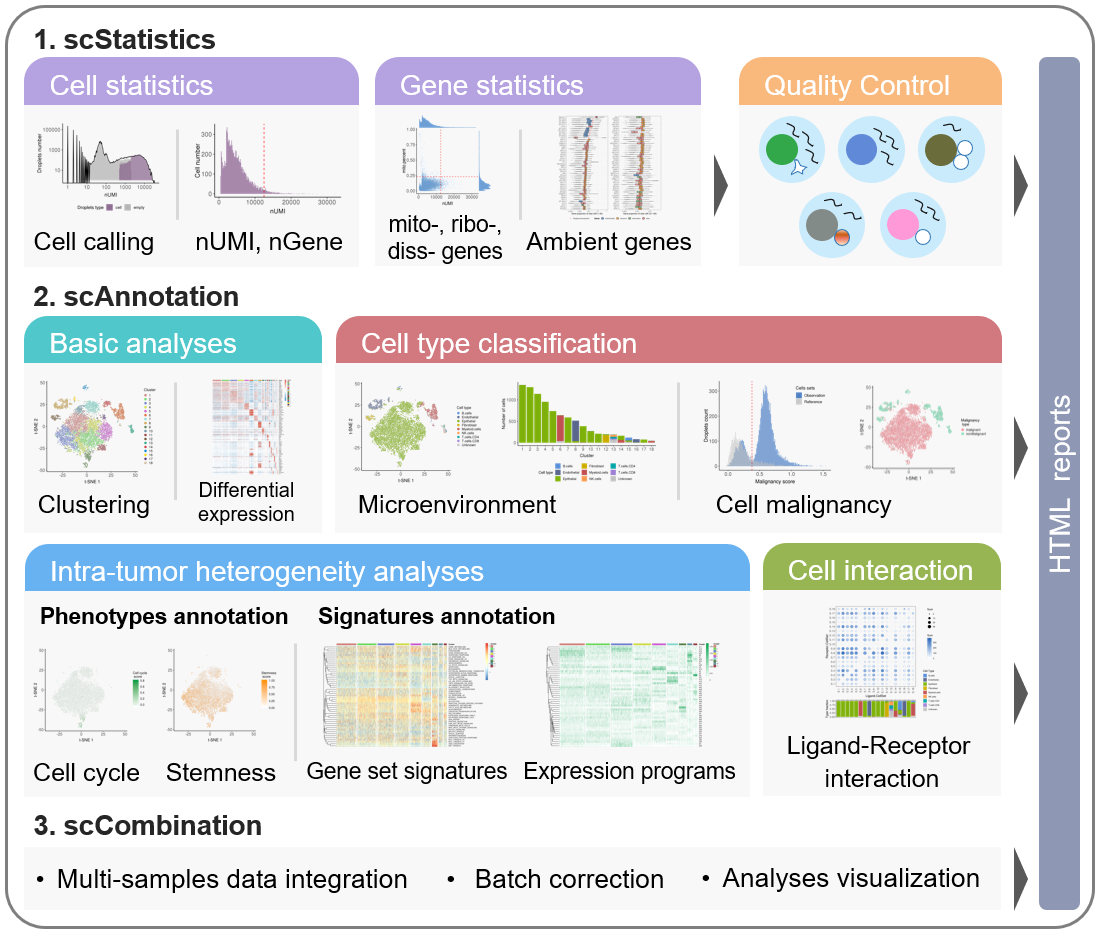
The workflow of scCancer mainly consists of three modules: scStatistics, scAnnotation, and scCombination.
-
The
scStatisticsperforms basic statistical analyses of raw data and quality control. -
The
scAnnotationperforms functional data analyses and visualizations, such as low dimensional representation, clustering, cell type classification, cell malignancy estimation, cellular phenotype analyses, gene signature analyses, cell-cell interaction analyses, etc. -
The
scCombinationperform multiple samples data integration, batch effect correction and analyses visualization.
After the computational analyses, detailed and graphical reports were generated in user-friendly HTML format.
- R version: >= 3.5.0
Firstly, please install or update the package devtools by running
install.packages("devtools")Then the scCancer can be installed via
library(devtools)
devtools::install_github("wguo-research/scCancer")Hint: A dependent package NNLM was removed from the CRAN repository recently, so an error about it may be reported during the installation.
If so, you can install a formerly available version manually from its archive.
Besides, if you encounter errors saying package SoupX is unavalibale, you can refer to its GitHub and install it via
devtools::install_github("constantAmateur/SoupX")library(scCancer)The scCancer is mainly designed for 10X Genomics platform,
and it requires a data folder containing the results generated by the software
Cell Ranger.
In general, the data folder needs to be organized as following which is the output of Cell Ranger V3:
/sampleFolder
├── filtered_feature_bc_matrix
│ ├── barcodes.tsv.gz
│ ├── features.tsv.gz
│ └── matrix.mtx.gz
├── raw_feature_bc_matrix
│ ├── barcodes.tsv.gz
│ ├── features.tsv.gz
│ └── matrix.mtx.gz
└── web_summary.html
Comparing to Cell Ranger V2 (CR2), Cell Ranger V3 (CR3) can identify cells with
low RNA content better. So we suggest to use CR3 to do alignment and cell-calling.
Considering that some published data is from CR2 or the raw matrix isn't supported, we
specially deisgn the pipeline to be compatible with these situations.
A common folder structure of CR2 is as below.
/sampleFolder
├── filtered_gene_bc_matrices
│ └── hg19
│ ├── barcodes.tsv
│ ├── genes.tsv
│ └── matrix.mtx
├── raw_gene_bc_matrices
│ └── hg19
│ ├── barcodes.tsv
│ ├── genes.tsv
│ └── matrix.mtx
└── web_summary.html
For other droplet-based platforms, the data folder should be prepared likewise.
Here, we provide an example data of
kidney cancer from 10X Genomics. Users can download it and run following scripts
to understand the workflow of scCancer. And following are the generated HTML reports:
For multi-samples, following is a generated HTML report for three kidney cancer samples integration analysis:
The scStatistics mainly implements quality control for the expression matrix
and returns some suggested thresholds to filter cells and genes.
Meanwhile, to evaluate the influence of ambient RNAs from lysed cells better,
this step also estimates the contamination fraction by using the algorithm of SoupX.
Following is the example script to run the first module scStatistics.
And using help(runScStatistics) can get more details about its arguments to realize personalized setting.
library(scCancer)
# A path containing the cell ranger processed data
dataPath <- "./data/KC-example"
# A path used to save the results files
savePath <- "./results/KC-example"
# The sample name
sampleName <- "KC-example"
# The author name or a string used to mark the report.
authorName <- "G-Lab@THU"
# Run scStatistics
stat.results <- runScStatistics(
dataPath = dataPath,
savePath = savePath,
sampleName = sampleName,
authorName = authorName
)Running the scStatistics script will generate some files/folders as below:
- report-scStat.html : A HTML report containing all results.
- report-scStat.md : A markdown report.
- figures/ : All figures generated during this module.
- report-figures/ : All figures presented in the HTML report.
- cellManifest-all.txt : The statistical results for all droplets.
- cell.QC.thres.txt : The suggested thresholds to filter poor-quality cells.
- geneManifest.txt : The statistical results for genes.
- ambientRNA-SoupX.txt : The results of estimating contamination fraction.
-
report-cellRanger.html : The summary report generated by
Cell Ranger.
Using the QC thresholds, the scAnnotation filters cells and genes firstly, and then
performs basic operations (normalization, log-transformation, highly variable genes identification,
unwanted variance removing, scaling, centering, dimension reduction, clustering,
and differential expression analysis) using R package Seurat V3.
Besides, scAnnotation also performs some cancer-specific analyses:
-
Doublet score estimation : In this step, we integrate two methods (binary classification based
bcdsand co-expression basedcxds) of R packagescdsto estimate doublet scores. -
Cancer micro-environmental cell type classification : In this step, we develop a data-driven OCLR (one-class logistic regression) model to predict cell types, including epithelial cells, endothelial cells, fibroblasts, and immune cells (CD4+ T cells, CD8+ T cells, B cells, nature killer cells, and myeloid cells).
-
Cell malignancy estimation : In this step, we refer to the algorithm of R package
infercnvto estimate an initial CNV profiles. Then, we take advantage of cells’ neighbor information to smooth CNV values and define the malignancy score as the mean of the squares of them. -
Cell cycle analysis : In this step, to analyze intra-tumor cell phenotype heterogeneity, we define cell cycle score as the relative average expression of a list of G2/M and S phase markers, by using the function “AddModuleScore” of
Seurat. -
Cell stemness analysis : In this step, to analyze intra-tumor cell phenotype heterogeneity, we define cell stemness score as the Spearman correlation coefficient between cells’ expression and our pre-trained stemness signature, by referring to the algorithm of
Malta et al. -
Gene set signature analysis : In this step, to analyze intra-tumor heterogeneity at gene sets level, we provide two methods to calculated gene set signature scores:
GSVAand relative average expression levels. By default, we use 50 hallmark gene sets fromMSigDB. -
Expression programs identification : In this step, to analyze intra-tumor heterogeneity at gene sets level, we use non-negative matrix factorization (NMF) to unsupervisedly identify potential expression program signatures.
-
Cell-cell interaction analyses : In this step, we referred to the methods of
Kumar et alto characterize ligand-receptor interactions across cell clusters.
Following is the example script to run the second module scAnnotation.
And using help(runScAnnotation) can get more details about its arguments to realize personalized setting.
library(scCancer)
# A path containing the cell ranger processed data
dataPath <- "./data/KC-example"
# A path containing the scStatistics results
statPath <- "./results/KC-example"
# A path used to save the results files
savePath <- "./results/KC-example"
# The sample name
sampleName <- "KC-example"
# The author name or a string used to mark the report.
authorName <- "G-Lab@THU"
# Run scAnnotation
anno.results <- runScAnnotation(
dataPath = dataPath,
statPath = statPath,
savePath = savePath,
authorName = authorName,
sampleName = sampleName,
geneSet.method = "average" # or "GSVA"
)Running the scAnnotation script will generate some files/folders as below:
- report-scAnno.html : A HTML report containing all results.
- report-scAnno.md : A markdown report.
- figures/ : All figures generated during this module.
- report-figures/ : All figures presented in the HTML report.
- geneManifest.txt : The annotation results of genes updated by filter information.
- expr.RDS : A Seurat object.
- diff.expr.genes/ : Differentially expressed genes information for all clusters.
- cellAnnotation.txt : The annotation results for each cells.
- malignancy/: All results of cell malignancy estimation.
- expr.programs/ : All results of expression programs identification.
- InteractionScore.txt : Cell clusters interactions scores.
The scCombination mainly performs multiple samples data integration, batch effect correction and analyses visualization based on the scAnnotation results of single sample. And four strategies (NormalMNN (default), SeuratMNN, Raw and Regression) to integrate data and correct batch effect are optional.
Following is the example script to run the module scCombination.
And using help(runScCombination) can get more details about its arguments.
library(scCancer)
# Paths containing the results of 'runScAnnotation' for each sample.
single.savePaths <- c("./results/KC1", "./results/KC2", "./results/KC3")
# Labels for all samples.
sampleNames <- c("KC1", "KC2", "KC3")
# A path used to save the results files
savePath <- "./results/KC123-comb"
# A label for the combined samples.
combName <- "KC123-comb"
# The author name or a string used to mark the report.
authorName <- "G-Lab@THU"
# The method to combine data.
comb.method <- "NormalMNN" # SeuratMNN Raw Regression
# Run scCombination
comb.results <- runScCombination(
single.savePaths = single.savePaths,
sampleNames = sampleNames,
savePath = savePath,
combName = combName,
authorName = authorName,
comb.method = comb.method
) Running the scAnnotation script will generate some files/folders as below:
- report-scAnnoComb.html : A HTML report containing all results.
- report-scAnnoComb.md : A markdown report.
- figures/ : All figures generated during this step.
- report-figures/ : All figures presented in the HTML report.
- expr.RDS : A Seurat object.
- diff.expr.genes/ : Differentially expressed genes information for all clusters.
- cellAnnotation.txt : The annotation results for each cells.
- expr.programs/ : All results of expression programs identification.
- (anchors.RDS : The anchors used for batch correction of "NormalMNN" or "SeuratMNN".)
If the sample data is generated by CR2 and contain "raw_gene_bc_matrices" matrix, the scCancer package can re-perform cell calling using the method EmptyDrop (the name of its R package is DropletUtils). But users need to manually install the DropletUtils and import it in script.
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("DropletUtils")
library(DropletUtils)After the step scStatistics, a HTML report (report-scStat.html) will be generated, which persents the statistical features of the data from various perspectives (nUMI, nGene, mito.percent, ribo.percent, diss.percent).
By identifying outliers from the distribution of these metrics, scCancer adaptively provides some suggested thresholds to filter low-quality cells and records them in the file cell.QC.thres.txt.
Users can modify the values in file cell.QC.thres.txt to make scAnnotation use the updated thresholds to perform cell QC and downstream analyses.
For the step gene QC, the scCancer filters genes according to three aspects of information.
- Mitochondrial, ribosomal, and dissociation-associated genes.
- Genes with the number of expressed cells less than argument
nCell.min(the default is 3). - Genes with the background percentage larger than the argument
bgPercent.max(the default is 1, which means unfilter). ThebgPercent.maxcan be decided according to the distribution of background percentage inreport-scStat.html.
Users can modify the values of these arguments according to their needs.
In the scStatistics module, users can pass in the soup (background) specific gene lists by the argument bg.spec.genes or use the default setting:
bg.spec.genes <- list(
igGenes = c('IGHA1','IGHA2','IGHG1','IGHG2','IGHG3','IGHG4','IGHD','IGHE','IGHM', 'IGLC1','IGLC2','IGLC3','IGLC4','IGLC5','IGLC6','IGLC7', 'IGKC', 'IGLL5', 'IGLL1'),
HLAGenes = c('HLA-DRA', 'HLA-DRB5', 'HLA-DRB1', 'HLA-DQA1', 'HLA-DQB1', 'HLA-DQB1', 'HLA-DQA2', 'HLA-DQB2', 'HLA-DPA1', 'HLA-DPB1'),
HBGenes = c("HBB","HBD","HBG1","HBG2", "HBE1","HBZ","HBM","HBA2", "HBA1","HBQ1")
)Then a contamination fraction will be estimated and saved in file ambientRNA-SoupX.txt.
In the scAnnotation module, if users want to correct the expression data according to the estimated ambient RNAs contamination fraction, they can set the argument bool.rmContamination as TRUE (the default is FALSE), and set the argument contamination.fraction as a number (between 0 and 1) or NULL (NULL means the result of scStatistics will be used).
By default, the arguments species and genome are set as human and hg19.
Users can set species as human or mouse, and set genome as hg19, hg38, or mm10, respectively.
Patient-drived tumor xenograft (PDX) samples contain both human and mouse cells generally. In order to deal with these human-mouse-mixed data, users can explicitly set the arguments hg.mm.mix, species, genome, hg.mm.thres, and mix.anno of the relevant functions.
More details for the meaning of these arguments can be found by using command help().
By default, we use both t-SNE and UMAP to obtain low-dimension coordiantes,
and the clustering results are presented using both of them.
For other analyses, we use t-SNE 2D coordinates by default.
Users can also set the argument coor.names as c("UMAP_1", "UMAP_2")
to view the results under UMAP coordinates.
Note: If users haven't installed UMAP, they can do so via
reticulate::py_install(packages = 'umap-learn')For the multi-modal data, which have both gene expression and antibody capture results, scCancer is also compatible. Users don't need to perform special setting, and scCancer will extract the expression data automatically and run downstream analyses.
If users' samples have other cell types to classify, they can train the new templates and pass in the argument ct.templates. To train the new templates, following cods can be referred:
library(gelnet)
train.data <- list("type1" = expr.data1, "type2" = expr.data2)
ct.templates <- lapply(names(train.data), FUN = function(x){
result <- gelnet(t(train.data[[x]]), NULL, 0, 1)
return(result$w[result$w != 0])
})Here is the output of sessionInfo() on the system.
R version 3.6.1 (2019-07-05)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: CentOS Linux 7 (Core)
Matrix products: default
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] scCancer_2.0.0
loaded via a namespace (and not attached):
[1] sn_1.5-4 plyr_1.8.5
[3] igraph_1.2.4.2 lazyeval_0.2.2
[5] GSEABase_1.48.0 splines_3.6.1
[7] BiocParallel_1.20.1 listenv_0.8.0
[9] GenomeInfoDb_1.22.0 ggplot2_3.2.1
[11] TH.data_1.0-10 digest_0.6.23
[13] htmltools_0.4.0 gdata_2.18.0
[15] magrittr_1.5 memoise_1.1.0
[17] cluster_2.1.0 ROCR_1.0-7
[19] globals_0.12.5 annotate_1.64.0
[21] matrixStats_0.55.0 RcppParallel_4.4.4
[23] R.utils_2.9.2 sandwich_2.5-1
[25] colorspace_1.4-1 blob_1.2.0
[27] rappdirs_0.3.1 ggrepel_0.8.1
[29] xfun_0.11 dplyr_0.8.3
[31] crayon_1.3.4 RCurl_1.95-4.12
[33] jsonlite_1.6 graph_1.64.0
[35] survival_3.1-8 zoo_1.8-7
[37] ape_5.3 glue_1.3.1
[39] gtable_0.3.0 zlibbioc_1.32.0
[41] XVector_0.26.0 leiden_0.3.2
[43] DelayedArray_0.12.1 future.apply_1.4.0
[45] SingleCellExperiment_1.8.0 BiocGenerics_0.32.0
[47] scales_1.1.0 pheatmap_1.0.12
[49] mvtnorm_1.0-12 DBI_1.1.0
[51] bibtex_0.4.2.2 miniUI_0.1.1.1
[53] Rcpp_1.0.3 metap_1.3
[55] plotrix_3.7-7 viridisLite_0.3.0
[57] xtable_1.8-4 reticulate_1.14
[59] bit_1.1-14 rsvd_1.0.2
[61] SDMTools_1.1-221.2 stats4_3.6.1
[63] tsne_0.1-3 GSVA_1.34.0
[65] NNLM_0.4.3 scds_1.2.0
[67] htmlwidgets_1.5.1 httr_1.4.1
[69] gplots_3.0.1.2 RColorBrewer_1.1-2
[71] TFisher_0.2.0 Seurat_3.1.2
[73] ica_1.0-2 pkgconfig_2.0.3
[75] XML_3.98-1.20 R.methodsS3_1.7.1
[77] uwot_0.1.5 tidyselect_1.0.0
[79] rlang_0.4.4 reshape2_1.4.3
[81] later_1.0.0 AnnotationDbi_1.48.0
[83] munsell_0.5.0 tools_3.6.1
[85] xgboost_0.90.0.2 RSQLite_2.1.5
[87] ggridges_0.5.2 stringr_1.4.0
[89] fastmap_1.0.1 npsurv_0.4-0
[91] knitr_1.26 bit64_0.9-7
[93] fitdistrplus_1.0-14 caTools_1.18.0
[95] purrr_0.3.3 RANN_2.6.1
[97] pbapply_1.4-2 future_1.16.0
[99] nlme_3.1-143 mime_0.8
[101] R.oo_1.23.0 ggExtra_0.9
[103] compiler_3.6.1 shinythemes_1.1.2
[105] plotly_4.9.1 png_0.1-7
[107] lsei_1.2-0 tibble_2.1.3
[109] geneplotter_1.64.0 stringi_1.4.5
[111] lattice_0.20-38 Matrix_1.2-18
[113] markdown_1.1 multtest_2.42.0
[115] vctrs_0.2.2 mutoss_0.1-12
[117] pillar_1.4.3 lifecycle_0.1.0
[119] Rdpack_0.11-1 lmtest_0.9-37
[121] RcppAnnoy_0.0.14 data.table_1.12.8
[123] cowplot_1.0.0 bitops_1.0-6
[125] irlba_2.3.3 gbRd_0.4-11
[127] GenomicRanges_1.38.0 httpuv_1.5.2
[129] R6_2.4.1 promises_1.1.0
[131] KernSmooth_2.23-16 gridExtra_2.3
[133] IRanges_2.20.1 codetools_0.2-16
[135] MASS_7.3-51.5 gtools_3.8.1
[137] assertthat_0.2.1 SummarizedExperiment_1.16.1
[139] sctransform_0.2.1 mnormt_1.5-5
[141] multcomp_1.4-12 S4Vectors_0.24.1
[143] GenomeInfoDbData_1.2.2 parallel_3.6.1
[145] SoupX_1.0.1 grid_3.6.1
[147] tidyr_1.0.2 Rtsne_0.15
[149] pROC_1.15.3 numDeriv_2016.8-1.1
[151] Biobase_2.46.0 shiny_1.4.0